Welcome to AFFIPred
AFFIPred is a web application for predicting pathogenicity of SNVs based on AlphaFold structures. AFFIPred provides predictions for all possible amino acid variations in human proteome.
AFFIPred is a web application for predicting pathogenicity of SNVs based on AlphaFold structures. AFFIPred provides predictions for all possible amino acid variations in human proteome.
Functional impact of missense variations has been extensively studied aiming to unravel sequential and/or structural patterns that would discriminate pathogenic variants from benign variants. Apart from sequence and structure-based features, protein dynamics has also been shown to play a role in pathogenicity of variations [1]. Protein structure, static or dynamic, contributes to our understanding of pathogenicity, albeit the pathogenicity classifiers, which rely on structural information, require an available structure in PDB. However, not all structures can be experimentally characterized nor all parts of a given protein chain could be resolved in a given structural study. To this end, AlphaFold succors to close the gap between the available sequence and structure data [2]. AlphaFold predicted structures have been recently deposited to PDB [3] comprising the largest fraction of the database. Nonetheless, AlphaFold structures represent predicted models that were not backed up by any empirical data contrary to the experimental structures. Thus, instead of blindly trusting the predicted structures, each structure should be assessed based on the confidence scores of the predicted position. This way, one can incorporate AF structures to understand structural patterns that underlie pathogenicity of missense variants.
1. Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic acids research. 2018;46(D1):D1062–D1067.
2. Consortium U. UniProt: a worldwide hub of protein knowledge. Nucleic acids research. 2019;47(D1):D506–D515.
3. Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nature methods. 2010;7(4):248–249
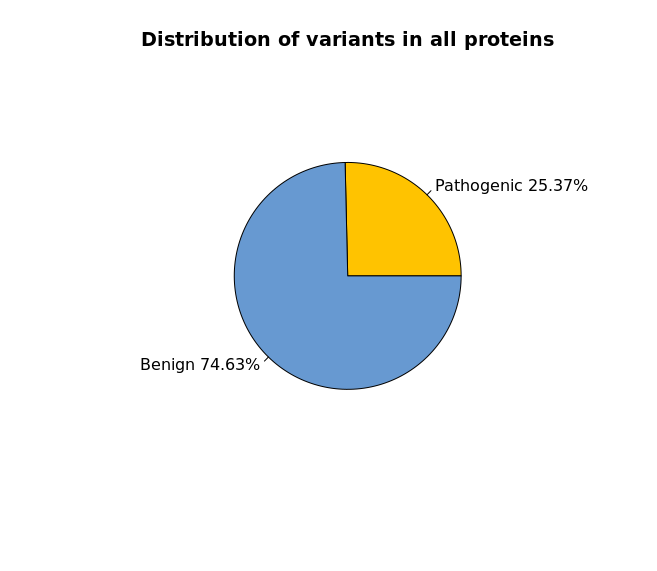
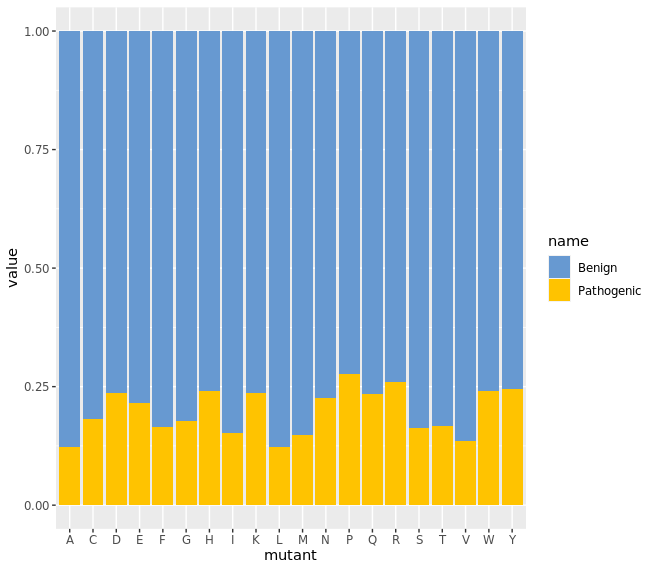
There are 2 main ways to query the database:
AF-based Functional Impact Prediction (AFFIPred) accepts multiple inputs including genomic position, amino acid position and variant number (RS id). Example query input for each format is available in the box.
Collect the necessary information about the variants you want to analyze. Organize this data in a format compatible with the AFFIPred’s requirements. Ensure that the prompt clearly specifies the gene/protein and variant information.
Genomic position: 19 1010539 G C
Protein position: P21817 Y116A
Variant ID: rs1042779
You can also upload a VCF file, or a txt file containing variants in one of these formats. If you upload a VCF file, you should select 'Genomic position' on the right panel.
Site saturation predictions can be visualized for a protein. Available input types are uniprot ID and gene name and Ensembl gene ID
Uniprot ID: Q04206
Gene name: TMEM145
Ensembl Gene ID: ENSG00000175899
First, install the tool using pip.
pip install -i https://test.pypi.org/simple/ affipred
Provide input and output files relative or absolute path:
affipred_pred variants.vcf -o affipred_results.csv